I am working with paired end fastq sequence data containing amplicons generated using several primers. I have tried two different methods to separate the amplicons by primer and to denoise but I am getting poor results.
For the first approach I used cutadapt to select the ZBJ amplicons and to cut off the ZBJ primers. The adapters were found in around 90% of the reads and 87% passed filtering.
qiime cutadapt trim-paired
--i-demultiplexed-sequences data.qza
--p-front-f AGATATTGGAACWTTATATTTTATTTTTGG
--p-adapter-f GGAGGATTTGGWAATTGATTAGTW
--p-front-r WACTAATCAATTWCCAAATCCTCC
--p-adapter-r CCAAAAATAAAATATAAWGTTCCAATATC
--p-match-read-wildcards
--p-match-adapter-wildcards
--p-discard-untrimmed
--o-trimmed-sequences ZBJ.qza
However, I don't think this was done correctly as the ZBJ amplicon should be around 157bp in length and the plot show my sequences are up to around 230bp.
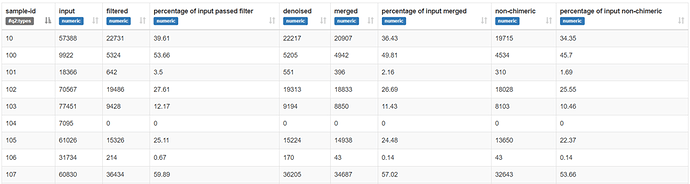
When I denoise these sequences with dada2 a high proportion of reads are removed during the initial filtering step. I have tried various dada2 settings including no trimming, trimming to 230bp and truncating from 130-220bp. Here is an example;
qiime dada2 denoise-paired
--i-demultiplexed-seqs ZBJ.qza
--p-trunc-len-f 150
--p-trunc-len-r 150
--p-trim-left-f 23
--p-trim-left-r 23
--p-n-threads 40
--o-table ZBJfeaturetable.qza
--o-representative-sequences ZBJrepseqs.qza
--o-denoising-stats ZBJstats.qza
As this gave poor results I decided to denoise the entire dataset.
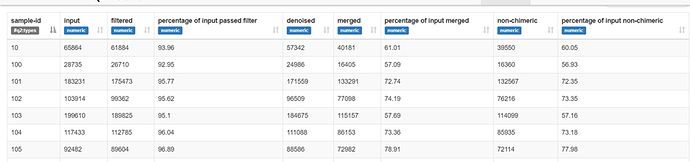
From this plot of all the sequences I used the following dada2 settings and got good results when truncating to 150bp seen in the stats table below (and poor results truncated to 160bp).
qiime dada2 denoise-paired
--i-demultiplexed-seqs dataimported.qza
--p-trunc-len-f 150
--p-trunc-len-r 150
--p-trim-left-f 23
--p-trim-left-r 23
--p-n-threads 20
--o-table allfeaturetable.qza
--o-representative-sequences allrepseqs.qza
--o-denoising-stats allstats.qza
As these results were good I thought I would try separating the amplicons following dada2 using qiime feature-classifier extract-reads as suggested here
qiime feature-classifier extract-reads
--i-sequences allrepseqs.qza
--p-f-primer AGATATTGGAACWTTATATTTTATTTTTGG
--p-r-primer WACTAATCAATTWCCAAATCCTCC
--p-n-jobs 20
--o-reads ZBJrepseqextracted.qza
However, this only gave a sequence count of 97 so I don't think the ZBJ amplicons were extracted correctly. Could this be due to the primers being degenerate? I changed the --p-n-jobs parameter to 0.7 but it had little impact.
So my questions are;
- What method should I use to separate my amplicons by primer?
- Why am I getting longer than expected sequence reads after using cutadapt?
- Why are so many sequences being filtered out during denoising after using cutadapt and what criteria does the first filtering step actually use?
- Can qiime feature-classifier extract-reads be used with degenerate primers?
Any clarification on these topics is greatly appreciated!