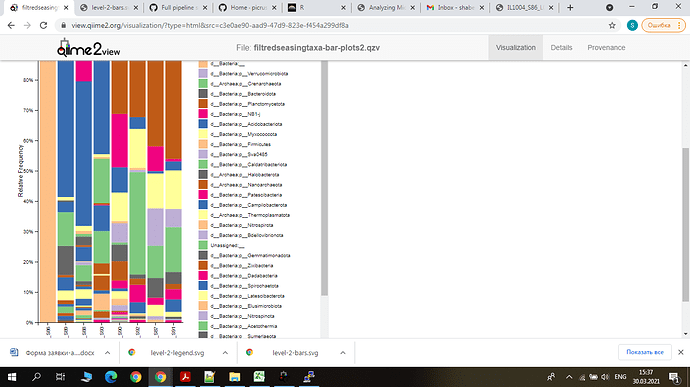
I have only one sample which contains only Bateria on the phylum level . What it would be? If there were Bacteria, why wasn't classified other levels?
Hi!
Could you provide more details, such as rDNA region amplified in your samples and which classifier you used, or how you trained your own classifier. For example, if you have V3-V4 region amplified, but used classifier for another region, you will receive a lot of unassigned or assigned only to bacteria features.
Salam Timur. Yes, I used the SILVA and analysed 16S rRNA by sequencing V3_V4 regions. Actually only in one from 8 samples showed that there are Bacteria. Did u say that it is unassigned? But here not indicated that they are unassigned. I meant I didn't find the word unassigned.
@ShonB, are you trying to understand why an expected taxon (Bacteria) is present in only one of your samples?
Alternately, are you trying to understand why Bacteria is the only piece of taxonomic information available in the annotation Bacteria;__;__;__.....?
Is it something else? Please take a few minutes to clarify your question - I'm not sure we have enough information to help.
Hi Chris.
Yes, I was wondering, why in my one of samples, is present only one taxa, Bacteria;__;__;__..... ? There not any other taxa on all levels. What it would be?
Sorry, @ShonB, but we still don't have enough information to help you troubleshoot this.
Maybe you can share the .qzv your screenshot is from? And some more details on your process? (what commands you ran, exactly what database and classifier you used, what you expected to see here, and why you suspect your result is incorrect)
Thanks for your patience!
Hi, Chris !
Thank you for reply. You are right, maybe I wasn't clear, so you couldn't understand me.
So, I was analysing 8 samples (only forward reads). And only in one sample nothing was assigned and it showed only Bacteria. I think it's impossible that in whole sample nothing assigned and there is only Bacteria. The reads are more 9800.
So, I was wondering, what I could do in this situation. Here, screenshot.
Thanks for clarifying the situation! Assuming those are similar samples, and they were treated similarly, I agree that that's very unusual.
I'd be happy to help you try to figure out why this happened. The best place to start is usually by looking at how you processed the Data. A full description is stored as "provenance" in every .qzv, so if you are able to share the actual taxa barplot file, you may not have to explain everything you did in detail.
Are there any steps in the analysis you were unsure about?
This topic was automatically closed 31 days after the last reply. New replies are no longer allowed.