Hi QIIME2 team,
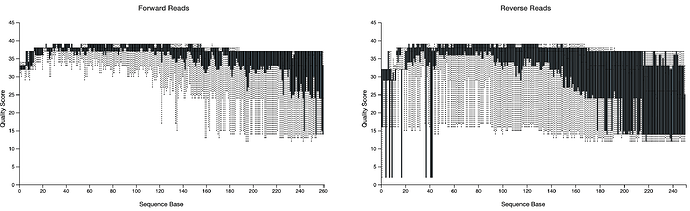
I have worked with QIIME2 to analyze 16S v4-5 data for several datasets and encountered an error I have never seen before in the steps from import, demux, to denoise. Because our lab recently also had some error code with read demultiplexing from sequenceing platform and I am just not sure where the error is acturally from. I have my qiime codes and errors copied below and the demux summary attached. My current suspicion is the reads number of 1/3 of my samples were too low, (<10,000 per sample ) so they failed denoise even if I set all the parameters to allow every reads imported.
Looking forward to your comments, thanks so much.
Best,
Shuqi
(qiime2-amplicon-2023.9) [host@compute-b-16-192 16s]$ cat qiime2.sbatch
#!/bin/bash
#SBATCH --job-name=qiime2
#SBATCH -p short
#SBATCH -t 0-12:00
#SBATCH --mem=64G
#SBATCH -c 16
#SBATCH -o %j.out
#SBATCH -e %j.err
module load conda/miniforge3/24.11.3-0
conda activate qiime2-amplicon-2023.9
cd 16s/
qiime tools import \
--type 'SampleData[PairedEndSequencesWithQuality]' \
--input-path 01-raw/ \
--output-path 02-demux/demux-paired-end.qza \
--input-format CasavaOneEightSingleLanePerSampleDirFmt
qiime demux summarize \
--i-data 02-demux/demux-paired-end.qza \
--o-visualization 02-demux/demux-paired-end.qzv
qiime dada2 denoise-paired
--p-n-threads 0
--i-demultiplexed-seqs 02-demux/demux-paired-end.qza
--p-trunc-len-f 0
--p-trunc-len-r 0
--o-representative-sequences 03-features/unfiltered-rep-seqs.qza
--o-table 03-features/unfiltered-table.qza
--o-denoising-stats 03-features/denoising-stats.qza
(qiime2-amplicon-2023.9) [host@compute-b-16-192 16s]$ cat err.log
Plugin error from dada2:
No reads passed the filter. trunc_len_f (0) or trunc_len_r (0) may be individually longer than read lengths, or trunc_len_f + trunc_len_r may be shorter than the length of the amplicon + 12 nucleotides (the length of the overlap). Alternatively, other arguments (such as max_ee or trunc_q) may be preventing reads from passing the filter.
Debug info has been saved to /tmp/qiime2-q2cli-err-85pe8y_a.log
P.S. I am working on remote cluster and have no access to the tmp file due to permission issue.