Can anyone help me with interpreting the dendogram tree that is produced using the qiime sample-classifier heatmap command? For reference, when I ran this command I left the clustering, etc. all in default. I tried looking at the seaborn documentation referenced in the documentation for the qiime sample-classifier heatmap command, but I am still not certain how to interpret the clustering. Any insight you could provide would be very much appreciated.
Hi @jrw187,
This dendrogram is the same one generated by qiime feature-table heatmap — the sample-classifier pipeline merely filters by important features and (optionally) groups the samples by category prior to generating the heatmap. So the interpretation is the same, though it's flavored by those caveats.
The dendrogram indicates the similarity between samples (or groups): samples (or groups) are clustered such that those next to each other are more similar than those that are far apart. The dendrogram shows this relationship and is interpreted like any other dendrogram, showing the hierarchical relationship between samples or groups of samples. Features can also be clustered by patterns of co-occurrence; features clustering together are more likely to co-occur, and those that do not cluster together are less likely to co-occur. In the case of q2-sample-classifier, the only nuance to this interpretation is that the features shown in the heatmap are the top N best predictors of whatever category you used.
The only real relevance of the dendrogram to q2-sample-classifier is that (assuming you did not group the samples) you would hope that samples belonging to the same group would mostly cluster together, demonstrating that relationship between that sample class and the predictive features.
You can view of this as a type of feature selection or "enrichment", whereby q2-sample-classifier finds the features that best differentiate your sample groups, then focuses on those in the heatmap, weeding out the "noise" created by all the other features. In the scenario that good prediction is observed, this will lead to clear clusters by sample class and obvious relationships between features and sample classes... your mileage may vary!
Thank you for your quick response.
I understand that the q2-sample-classifier (if your prediction model accuracy is good) should give you a sense of positive/negative responders and co-occurrence of features, so very valuable tool. Thank you for a very clear explanation of this.
I am going to attempt to ask you a follow up question here (bear with me, I am less clear on how to ask this and what exactly I am asking for, but I am going to give it a shot). I am fairly new this type of analysis and have been working with a collaborator whose lab is totally focused on microbiome research. Their lab only really uses Qiime2 to generate the feature table and assign taxonomy and then they move over a couple of different R packages for their analysis, so I have been getting a lot of questions about interpreting the output from q2-sample-classifier (which is good because it is pushing me to really get a better understanding of the analytical methods I am using). The latest question has been about interpreting the tree. Obviously it is hierarchical and obviously it is indicative of potential co-occurence (as you so well-described in your previous answer). But I think what my collaborator is asking about it how that clustering is done - what are the parameters. In the documentation it indicates that by default the clustering metric is bray-curtis, the method is average, and it clusters by feature. Do I interpret this to mean that the command clusters features based on the average bray-curtis distance? What kind of coordinates is it using to calculate the bray-curtis distance (not sure if this is the correct way to ask this!)? Any clarification that you can provide on this would be helpful - would love to have a bit more detail to provide in my next meeting regarding this topic!
Thanks!
For reference, I am grouping by a categorical variable when I do this and I have seen high accuracy in the prediction modeling for some datasets and not-so-good for others. The dataset that sparked this conversation was one where the model prediction was good, so there is some obvious clustering.
oh got it, thanks for clarifying your question!
Yes
It is creating a bray-curtis distance matrix based on abundance of input features... so in this case it is bray-curtis distance between samples based on abundance of only these top most important features. And bray-curtis distances between features based on their abundances across samples.
Great, nothing like a well-clustered heatmap!
Hello @Nicholas_Bokulich ,
I have one additional follow-up question on this. Can you tell me anything about the scaling of the branch lengths on the heatmap?
My collaborator is interested in seeing a scale bar to indicate the value/meaning of the branch lengths.
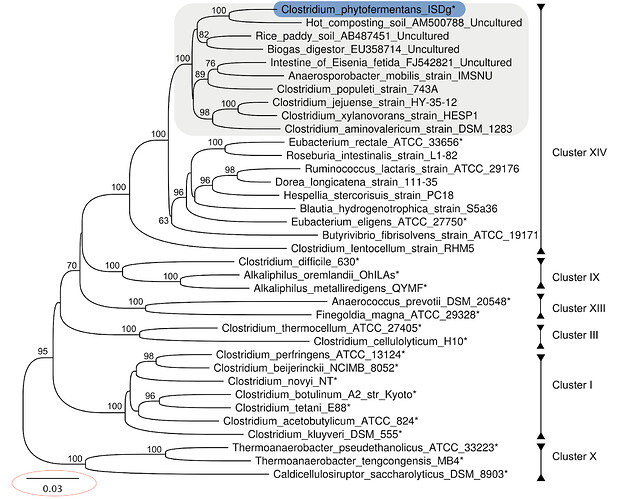
Below is a screenshot with the notation circled so you can see what I am talking about. Not the same kind of tree, but that line indicates the difference between the sequences in each cluster.
Thank you again for all of your help with this.
No, sorry — you would need to check the seaborn documentation for that info
This topic was automatically closed 31 days after the last reply. New replies are no longer allowed.