Hello,
I am trying to import a manifest file but keep getting an error.
Here is what I am running:
qiime2) dh103@8BFS8B3-LINUX:~$ qiime tools import --type 'SampleData[SequencesWithQuality]' --input-path /home/dh103/Documents/Greta/Cleandata_Manifest2.0.csv --output-path CleanData_demux.qza --input-format SingleEndFastqManifestPhred33V2
and this is the error i am receiving:
Traceback (most recent call last):
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/q2cli/builtin/tools.py", line 157, in import_data
artifact = qiime2.sdk.Artifact.import_data(type, input_path,
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/sdk/result.py", line 277, in import_data
return cls.from_view(type, view, view_type, provenance_capture,
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/sdk/result.py", line 305, in _from_view
result = transformation(view, validate_level)
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/core/transform.py", line 68, in transformation
self.validate(view, level=validate_level)
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/core/transform.py", line 143, in validate
view.validate(level)
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/plugin/model/file_format.py", line 26, in validate
self.validate(level)
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/q2_types/per_sample_sequences/_format.py", line 40, in validate
md = qiime2.Metadata.load(str(self))
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/metadata/metadata.py", line 357, in load
return MetadataReader(filepath).read(into=cls,
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/metadata/io.py", line 71, in read
header = self._read_header()
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/metadata/io.py", line 130, in _read_header
for row in self._reader:
File "/home/dh103/anaconda3/envs/qiime2/lib/python3.8/site-packages/qiime2/metadata/io.py", line 69, in
self._reader = (self._strip_cell_whitespace(row)
_csv.Error: ' ' expected after '"'
An unexpected error has occurred:
' ' expected after '"'
See above for debug info.
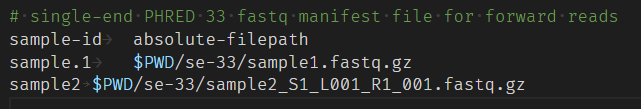
I believe qiime2 wants a tab after " but I have not used " when creating my manifest file so I am unsure how to fix this error.